©Copyright 2018 GEOSCIENCE RESEARCH INSTITUTE
11060 Campus Street • Loma Linda, California 92350 • 909-558-4548

A CREATIONIST VIEW OF CHROMOSOME BANDING AND EVOLUTION
by
L. James Gibson
Geoscience Research Institute
WHAT THIS ARTICLE IS ABOUT
Similarities in organisms are commonly interpreted as the result of a common ancestry. Since chromosomes are the carriers of heredity, similarities in chromosomes could have special significance in studying the ancestry and relationships of species. Many studies comparing chromosomal banding patterns have been conducted. Through the use of a special staining technique, chromosomes can be stained to show a pattern of alternating dark and light bands. The detail of the pattern depends on the length of the chromosome at the time it is stained. If sufficient detail is present, each pair of chromosomes in a cell can be distinguished. Chromosomes from different species can be stained and then compared and contrasted. Similarities and differences maybe interpreted subsequently as reflecting the degree of relationship.
Although there are some interesting exceptions, comparisons of chromosomal banding patterns are generally consistent with comparisons based on other criteria. Species within a family generally show considerable matching of chromosome banding patterns. The cat, camel, and cow families each have substantial intrafamilial similarities in chromosomal banding patterns. On the other hand, some species have banding patterns which differ greatly from those of other species in the same genus. Similar species with different chromosomal banding patterns are found among certain deer (the muntjacs) and bats (Family Phyllostomidae).
More problematic are the similarities in chromosomal banding patterns among species which are different in structure. Interfamilial chromosomal banding similarities are found among the cats, mongooses, and raccoons; among the cow, deer, and giraffe families; among several families of marsupials; and among several families of primates, including humans. This raises questions about the extent of change which may have occurred in mammals, as well as the relationship of humans to other primates.
Four hypotheses to explain similarities of chromosomal banding are discussed in this paper. Such similarities could be the result of common design, of common ancestry, of chance, or of the action of virus-like agents. The hypothesis that chromosomal similarities could be due to chance seems unreasonable, It seems more likely that virus-like agents would cause differences between karyotypes than that they would change different karyotypes to look similar. Common ancestry appears to be the most likely basis for chromosomal similarities in species classified in the same genus, and for some species classified in different genera. However, to extend this explanation to higher taxonomic categories, in which similarities are of lesser extent and of lower quality does not seem necessary. To a creationist, it seems more probable that chromosomal similarities such as are found within the artiodactyls, the carnivores, the marsupials or the primates may be the result of common design.
Do different species of mammals have similar chromosomes? What is the meaning of chromosomal similarities? Are they evidence of common ancestry? Do they provide any support for common design? This article will discuss some of the evidence and attempt to propose answers to these questions.
INTRODUCTION
The genetic instructions for an organism are located in the chromosomes of the cells of the organism and are transmitted to the offspring by inheritance. A logical prediction of evolutionary theory is that closely related species should have similar chromosomes. Techniques of chromosome banding have now been available for a long enough period of time that some trends have been discovered, and the results can be examined profitably.
Comparisons of karyotypes (sets of chromosomes) can be based upon differing levels of detail (see White 1978:47). The first comparisons were made on the basis of the number of chromosomes. In some cases the number of one-armed (acrocentric) and two-armed (metacentric) chromosomes were included in the comparison, and the sex chromosomes were identified. Extensive lists of chromosome counts can be found in Matthey (1973a,b) for placental mammals and in Sharman (1973) and Hayman (1977) for marsupials. However, attempts to infer relationships based upon unbanded karyotypes have not been satisfactory (Atchley 1972). Frequently, individual chromosomes could not be identified, making comparisons of uncertain validity. Differences in arm number due to gain or loss of heterochromatin (tightly condensed chromatin, generally considered to have little genetic activity) were not correctly interpreted using conventional staining (Duffy 1972).
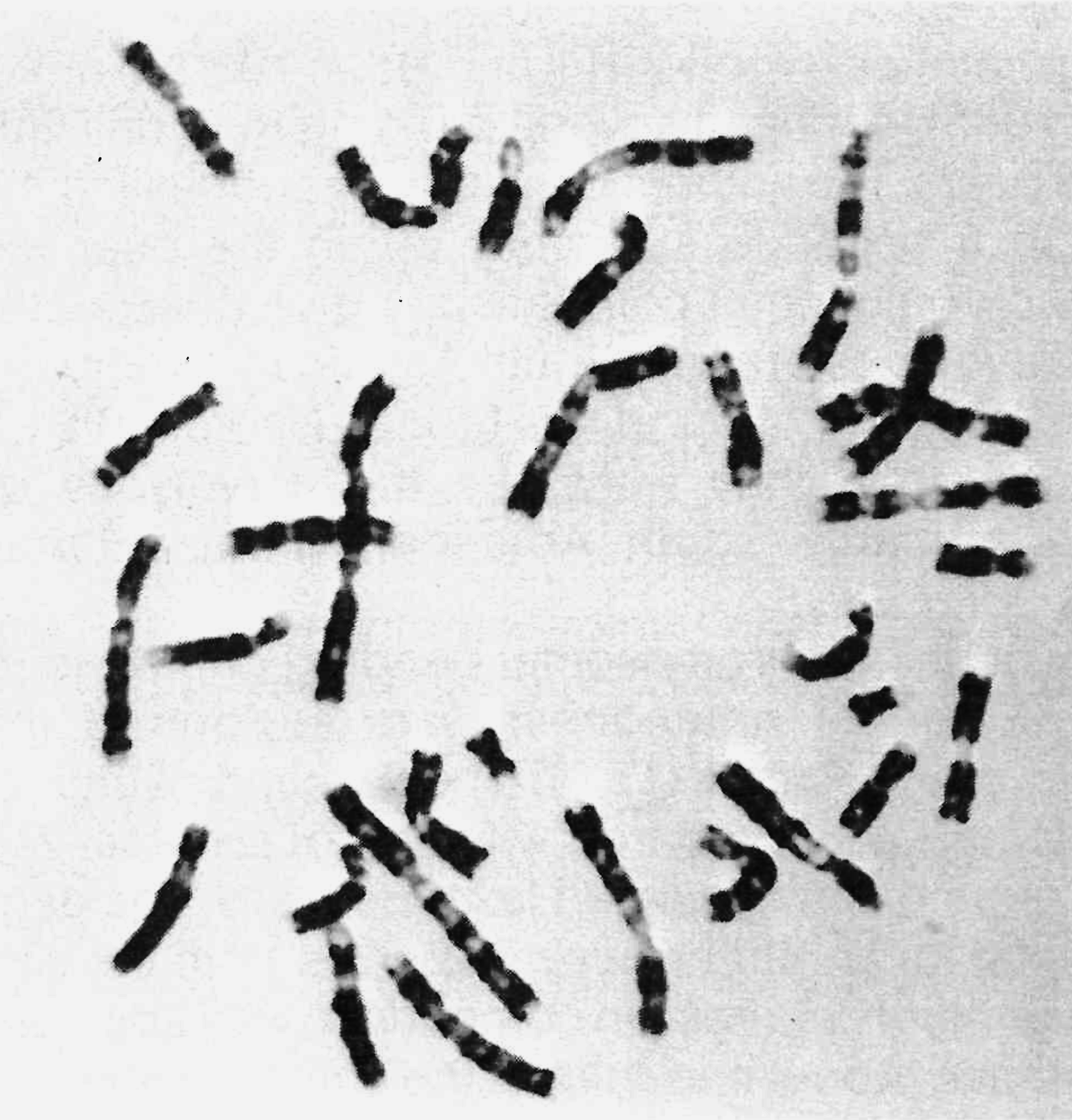
The development of banding techniques overcame these difficulties and made comparisons more meaningful. Structural changes in chromosomes (chromosomal rearrangements) can now be identified precisely. However, much remains to be learned about the meaning of banding and the structure of chromatin (the chromosomal material), and further developments can be expected to add to the value of comparative karyology. Several methods of chromosomal banding are available, but the most widely used method is G-banding (Giemsa-banding). This technique produces a characteristic pattern of contrasting dark and light transverse bands on the chromosomes (see Figure 1). The banding pattern is different for nearly all species studied, although sometimes the differences are slight. A large number of mammal species have been G-banded, but the number of species remaining to be studied is much larger.
PURPOSES OF COMPARISONS
Homeology
Comparisons of banding patterns often reveal nearly identical patterns in closely related species. The corresponding bands are believed to be homologous, but to allow for minor genetic difference, the term "homeology" is often used (e.g., Dutrillaux, Couturier and Fosse 1980).
Chromosomal similarities have been noted between groups of species in different genera or higher taxonomic categories. A significant degree of homeology has been found among all three families of seals (Arnason 1977). Homeologies at the suborder level have been noted within the primates (e.g., Dutrillaux, Couturier and Fosse 1980). Comparison of cat and human banding patterns (Nash and O'Brien 1982) did not show significant banding homeologies, although their gene mapping studies suggest similar gene arrangements (O'Brien and Nash 1982). Claims of banding homeologies between primates and carnivores by Dutrillaux and Couturier (1983) are of uncertain validity, because the proposed homeologies are based on portions of chromosome arms with a small number of bands involved. Reliable homeologies should include entire arms or be supported by other evidence (see Ponsa et al. 1981).
The actual genetic homology of similar banding patterns is supported by comparative gene mapping. Genes for equivalent enzymes are indeed often present on chromosomes with similar banding patterns in different species (Lalley and McKusick 1985). Genes which are found close together on a chromosome are said to be linked. Groups of genes which are linked in humans are generally also found linked in other species. In fact, gene groupings appear to be similar in many species even when chromosome banding homeology has not been detectable (Nash and O'Brien 1982, Kiel et al. 1985). Several equivalent linkage groups have been found on chromosomes with similar banding patterns in humans and other primates (Lalley and McKusick, loc. cit.). Several linkage groups are also common to human and cat chromosomes; in fact, the similarities in linkage patterns between cats and humans are almost as consistent as between chimpanzees and humans (O'Brien and Nash 1982), although based on fewer gene loci. Several mouse linkage groups are similar to human linkage groups, if one allows for the correspondence of one two-armed human chromosome with two one-armed mouse chromosomes. Whether the genes controlling such characteristics of organisms as development and morphology are also linked in similar ways in similar species is not known.
Classification
Comparative studies of chromosomal banding patterns have been useful in classification. Sibling species are species which appear alike morphologically, but have been discovered to be reproductively isolated. Giemsa-banding is not necessary to detect sibling species, but it can assist in identifying the differences between the species more precisely. Chromosomal sibling species have been discovered within cotton rats (Sigmodon; see Elder 1980), grasshopper mice (Onychomys; Hinesley 1979), and shrews (Sorex; Olert and Schmid 1978).
The use of G-banding has sometimes been helpful in clarifying the taxonomic position of species which do not have clear affinities based on other characters. Chromosomal differences have been used in determining the taxonomic placement of the African rat Mastomys (Lee and Martin 1980), the golden mouse (Ochrotomys; Engstrom and Bickham 1982), and Neotomodon alstoni (Yates, Baker and Barnett 1979). The giant panda, Ailuropoda, has been variously classified with the bears, raccoons, or in a family by itself. Although previous researchers were unable to find good banding matches (Wurster-Hill and Bush 1980), a more recent study (O'Brien et al. 1985) identified several banding matches linking the giant panda with the bears. The authors conclude by suggesting the giant panda be classified in a separate subfamily within the bear family Ursidae.
Sometimes chromosomal comparisons give unexpected results. For example, the karyotypes of some South American genera of rodents were more like the karyotypes of wood rats (genus Neotoma) a primarily North American genus, than they were like the karyotypes of cotton rats (Sigmodon), a primarily South American group (Baker, Koop and Haiduk 1983).
The validity of using chromosomal banding comparisons to assist in determining degree of relatedness was supported in a study by Mascarello, Stock and Pathak (1974). The banding pattern of a species of woodrat (Neotoma) was compared with that of six other rodent species, progressively more distantly related taxonomically. The degree of matching was near total for another woodrat species from the same subgenus and showed a general decrease in comparison with species of increasing taxonomic distance. About one-third of the chromosomes matched those of species from other tribes or subfamilies. Comparison with a species of kangaroo rat (Dipodomys, different superfamily) revealed no detectable banding homeologies with the woodrat. These results show general agreement with traditional methods of classification.
Constructing phylogenies
Giemsa banding has made possible the identification of specific chromosomes involved in rearrangements (Seabright 1972), permitting one to determine whether similar species possess the same rearrangement. Shared chromosomal rearrangements in similar species are interpreted as evidence of a common ancestor which had the rearrangement (see Rofe 1976). Several species of antelopes which share a Y/autosome translocation provide one such example (Benirschke et al. 1980).
The ability to identify similar banding patterns in different species and to identify chromosomal rearrangements has led to interest in reconstructing the historical sequence of rearrangements which have accompanied speciation in a group of species (see Spotorno 1977). The construction of such "family trees" is based on several assumptions. One assumption is that the ancestral karyotype can be determined with reasonable accuracy. This requires that the species to be compared be chosen carefully (Dutrillaux and Couturier 1983) and that homeologies be accurately identified. Another assumption is that the best tree is one which requires the fewest reversals and convergences (principle of parsimony; see Farris 1978).
The method is not without difficulties. One problem is that reversals and convergences do occur (e.g., see Baker, Barnett and Greenbraum 1979; Baker, Koop and Haiduk 1983; Searle 1984), probably because certain points on a chromosome are more susceptible to breakage than other points (Bush 1981, Nevers and Saedler 1977). Another problem is the possibility of mismatches, especially when only portions of chromosomal arms are involved (see Ponsa et al. 1981). Despite these difficulties, cladograms ("family trees") based on chromosomal characters are useful in testing for congruence with cladograms based on other data.
Cladograms based on chromosomal characters have been constructed for groups at several taxonomic levels, for example the genus Peromyscus (Robbins and Baker 1981), the bovid tribe Tragelephini (Benirschke et al. 1980), and several genera of murid and cricetid rodents (Koop et al. 1984). An especially comprehensive study at the superfamily level has been done for phyllostomoid bats (Patton and Baker 1978). Obviously, such cladograms cannot be more accurate than the identification of banding homeologies.
RESULTS OF COMPARATIVE G-BANDING STUDIES
This section discusses numerous examples of studies in which banding patterns of various species have been compared. It forms the basis for the discussion in the next section (A creationist viewpoint). Readers not interested in details may skip to the next section, referring to this section only if more details are desired. No attempt is made here to present an exhaustive list of references on G-banding results in mammals. Instead, I will discuss briefly a sample of the literature available, emphasizing studies of special interest or taxonomic breadth. Preference is given to papers which compare the banding patterns among several species or higher categories.
Monotremes
The taxonomic relationships of the egg-laying mammals are somewhat uncertain, as they show some skeletal similarities with reptiles and others with mammals (see Nowak and Paradiso 1983:1). There are three living genera, divided into two families. The anatomical uniqueness of the species is paralleled by the unusual nature of the karyotypes. The platypus has 52 chromosomes in each sex (Bick and Sharman 1975). Like most mammals, males have an XY sex chromosome pair, and females have two X chromosomes. Tachyglossus, the more widespread genus of echidna, is the only monotreme for which G-banding has been published (Murtagh 1977). There are 63 chromosomes in the male, which has one Y and two X chromosomes (X1X2Y). The female has two pairs of X chromosomes (X1X1X2X2) with 64 chromosomes in all. Zaglossus, the other genus of echidna, apparently has the same system. Unlike most other mammals, all of the monotreme species studied have unpaired elements at meiosis, which participate in a chain multiple with the sex chromosomes. The three genera of monotremes are karyotypically more similar to each other than to any other mammal.
Marsupials
The marsupials are of special interest because of their unusual characteristics and biogeographic distribution. Chromosome numbers for over 100 species of marsupials are listed by Hayman (1977), and trends analyzed. The chromosome numbers range from 10 to 32, with 14 being the most frequent number, and 22 the next most frequent. The greatest diversity of chromosome number is among the Macropodidae (kangaroos etc.).
Comparing Australian and American marsupials. The relationship of Australian marsupials to those from South America is a question of continuing interest. An important interfamilial comparison of G-banding by Rofe and Hayman (1985) may shed some light on the question. The study included one American species and 14 Australian species, representing four or five superfamilies (depending on the classification scheme). All species had 14 chromosomes, and their banding patterns showed remarkable agreement. These results were interpreted by the author as supporting the common ancestry of both Australian and American marsupials, with the ancestral karyotype being most like the one shared by a wombat (Vombatus ursinus), a dormouse possum (Cercartetus concinnus), and a bandicoot (Isoodon obesulus). Differences between species can be accounted for on the basis of pericentric inversions (an inversion including the centromere) and small variations in heterochromatin. The presumed increase in 2n number from the proposed ancestral number of 14 to as many as 32 is attributed to chromosomal fission.
The South American marsupial Dromiciops has sometimes been placed in a family separate from other living species. Its chromosomes have not been G-banded, but they appear to be similar to those of the presumed ancestral marsupial 2n=14 karyotype (see Sharman 1982). Karyotypic similarities have also been reported among several species of American opossums (Yonenaga-Yassuda et al. 1982; Casartelli, Rogatto and Ferrari 1986), and among several species of Australian dasyurid marsupials (Young et al. 1982, Baverstock et al. 1983b, Rofe and Hayman 1985).
Kangaroos. The chromosomes of kangaroos and their allies appear to be distinct from those of other marsupials. Rofe (1976) compared the G-bands of ten species of kangaroos and their allies, with 2n ranging from 10 to 22. Numerous chromosome arm homeologies could be identified, suggesting Robertsonian fusion (fusion of two chromosomes by their centromeres) to be the predominant type of chromosome rearrangement. The karyotype of the red-bellied pademelon (Thylogale billardierii), with 22 chromosomes, was interpreted as being closest to the ancestral condition for the group. Karyotypes for seven species of kangaroos (Macropus) were derivable by various fusions. The banding patterns of the swamp wallaby (Wallabia bicolor: 10 chromosomes in males, 11 in females) and the rock wallaby (Petrogale penicillata: 22 chromosomes, but a different karyotype) displayed greater divergence from the presumed ancestral state. Pericentric inversions and centric shifts (change in position of the centromere) were proposed to explain the differences.
Insectivores
One of the first examples of chromosomal polymorphism to be discovered was the European shrew superspecies, Sorex araneus. At least 12 chromosomal forms have been described (see Seale 1984). Differences can be explained on the basis of Robertsonian fusions involving different acrocentrics of an ancestral karyotype. Chromosome numbers range from 20 to 32. Some hybridization occurs, but partial reproductive isolation exists between some of the races. An X-autosome fusion, giving rise to a sex chromosome system of XX in females and XY1Y2 in males, is found in some, but not all of the races (Olert and Schmid 1978).
Bats
Bats comprise the second-most-diverse order of mammals. Two suborders are present, the Old World fruit bats and the rest of the bats. Eight species of African fruit bats, representing eight genera, were compared on the basis of their G-bands (Haiduk et al. 1981). In spite of the fact that chromosome numbers ranged only from 34 to 36, substantial differences in banding patterns were detected, requiring at least 34 rearrangements to explain the differences among the eight species. The mechanism for nearly half the rearrangements could not be conclusively identified. This study illustrates that significant karyotypic differences may exist between karyotypes which appear similar superficially.
Most G-banding studies of bats are concerned with the insectivorous and nectar-feeding bats. Patton and Baker (1978) concluded that the ancestral karyotype for the largely tropical American superfamily Phyllostomoidea is most like that of the big-eared bat (Macrotus waterhousii). In two different genera, a comparison of banding patterns of two similar species suggested that a total rearrangement of the genome had occurred in one species but not in the other. Another interesting discovery in this study was that the fisherman bat (Noctilio), placed in its own family on morphological grounds, has very similar G-banding to that of the mustache bat (Pteronotus pamellii), family Mormoopidae. The existence of similar karyotypes in morphologically distinct species and of different karyotypes in morphologically similar species can be interpreted as evidence against the theory that chromosomal rearrangements promote speciation by disruption of genes which regulate development (e.g., Wilson, Sarich and Maxson 1974).
Other good studies of chromosomal banding in bats include those of Haiduk and Baker (1982) on the long-tongued bats (Glossophaginae) and on evening bats by Bickham (1979a,b) and Zima (1982). A summary of the kinds of chromosomal rearrangements proposed in various studies of New World bats was published by Baker and Bickham (1980).
Primates
Because of the great interest in their relationship to humans, primates have been the object of special attention in comparative karyology. Numerous banding homeologies have been claimed for some 60 species of primates, including man (Dutrillaux et al. 1978). They conclude that "it is likely that all the euchromatin [genetically active chromatin] ... is identical in all the species". This statement, if true, would appear to reduce the significance of chromosomal banding comparisons.
As is often the case, different types of chromosomal rearrangements are typical of different taxonomic groups (karyotypic orthoselection; White 1973). In most lemurs Robertsonian rearrangements are the most common type of rearrangement, except for one genus in which tandem fusions are common (Rumpler and Dutrillaux 1976, 1978, 1979; Rumpler et al. 1983b, 1985). If their karyotypes are derived from the presumed ancestral karyotype for the group, Robertsonian rearrangements predominate in the species of Galago, while pericentric inversions are more important in Perodicticus, a loris (Dutrillaux et al. 1982, Rumpler et al. 1983a). Chromosome fissions are reportedly very frequent in the Old World monkeys, but have not been found in the other families. In the apes pericentric inversions are the most common type of rearrangement.
One of the most variable genera karyotypically is the New World owl monkey, Aotus. Nine different karyotypes have been reported, differing by fissions, fusions, and inversions (Ma 1981, Galbreath 1983). The number of sex chromosomes differs among the races. An ancestral karyotype for platyrrhine (New World) monkeys was proposed by Dutrillaux and Couturier (1981). A bibliography of cytogenetic studies in New World primates is available (Mudry de Pargament, Brieux de Salum and Colillas 1984).
Different workers have sometimes obtained different results from study of the same material. This can be illustrated in studies comparing the banding patterns of the grivet (Cercopithecus aethiops) and the rhesus monkey (Macaca mulatta). One pair of investigators (Stock and Hsu 1973) reported complete matching of the euchromatin (genetically active chromatin), with differences explainable as the result of heterochromatin additions or fusions. Another group of investigators (Estop, Garver and Pearson 1978) were unable to match some of the chromosomes in the two species. The claim of nearly complete homeology of banding among the Old World monkeys (Dutrillaux 1979, Dutrillaux et al. 1978) has been questioned by Ponsa et al. (1981), who suggest that the extent of banding homeologies among the primates may have been overstated. They emphasize the need to base homeologies on characteristic banding patterns, not merely short segments, and criticize the construction of karyotypes of hypothetical ancestors as "paper cytology".
Apes and humans. The chromosomes of the great apes have received a great deal of study, and detailed banding patterns have been published and compared with human banding patterns (Yunis and Prakesh 1982). The similarities between chimpanzee and human chromosomes are very striking. Only ten of the 23 pairs of human chromosomes show banding differences when compared with chimpanzee chromosomes. The banding patterns of nine chromosomes are identical in humans and gorillas. The three species differ in their banding patterns by various inversions and a Robertsonian fusion. The fusion involves chimpanzee chromosomes 12 and 13 as equivalent to human chromosome 2 (see Sun, Sun and Ho 1978a,b). No differences in the gene maps of humans and chimps have yet been noted (Lalley and McKusick 1985). In contrast to the usual phylogeny proposed for the group, it is the human karyotype that is considered to be closest to the ancestral type (Yunis and Prakesh 1982). The karyotypes of chimps and gorillas are more similar to the human karyotype than to that of the orangutan.
Carnivores
Banding similarities among the cat, raccoon and mongoose families were reported by Wurster-Hill and Gray (1975). More recently, attempts have been made to propose an ancestral karyotype for the order Carnivora (Dutrillaux and Couturier 1983, Couturier et al. 1986). This hypothetical karyotype is quite similar to that of the palm civet (Paradoxurus hermaphroditus, family Viverridae). Seal karyotypes show banding similarities with those of the carnivore families, but bears and dogs have karyotypes that are quite different.
The hypothetical ancestral carnivore karyotype (see above paragraph) was compared with the hypothetical ancestral karyotype previously proposed for the New World monkeys (Dutrillaux and Couturier 1981), prosimian primates (Rumpler et al. 1983b) and the squirrels (Petit et al. 1984, cited by Couturier et al. 1986). Although banding homeologies are claimed for significant portions of the karyotype, the method used has been criticized (Ponsa et al. 1981). Dutrillaux and Couturier invoke gene mapping similarities to support their view of actual homology of the chromosomes.
Karyotypically, one of the most homogeneous families known is the cat family (Wurster-Hill and Gray 1973). Mongooses show significant but varying degrees of similarity with cats (Wurster-Hill and Gray 1975). Translocations involving a sex chromosome are known in at least two genera of mongooses (Pathak and Stock 1976, Fredga 1972).
Seals and their allies
There are three families of pinnipeds: true (earless) seals, sea lions (eared seals), and the walrus. These families all share considerable banding homeology, with only four different karyotypes known (Arnason 1977). Differences among the karyotypes were not described thoroughly, but at least one fusion is involved. A striking resemblance to certain carnivore karyotypes was reported, especially to the coati mundi karyotype, but it is not clear whether this similarity was based on banding patterns.
Whales
Whales are generally divided into two major groups, toothed whales (Odontoceti) and baleen whales (Mysticeti). Karyotypes of members of both groups are very similar (Arnason 1974, Arnason et al. 1977), except for the sperm whales, which have distinctive karyotypes. Several species have interstitial heterochromatin and similar C-bands. Some homeologies were reported, but differences have not been described.
Odd-toed ungulates
Horses are the only members of this order for which I have seen comparative G-banding studies. All seven living species of the horse family have been studied (Ryder, Epel and Benirschke 1978). Each species has a different 2n number, ranging from 32 to 66. Only the X chromosome and a single autosome show the same banding pattern in each species. The other chromosomes all show differences, most commonly involving Robertsonian fusions and pericentric inversions. The mechanism for many of the rearrangements is unknown. The two species of horses have similar chromosomal banding patterns, as do the two species of asses. Two of the three species of zebras have similar patterns, but the pattern in Hartman's zebra is so different that little homeology can be determined in comparisons with the other species.
Even-toed ungulates
Interfamilial G-band homeologies have been identified (Buckland and Evans 1978) among the cow family (Bovidae), deer family (Cervidae) and the giraffe family (Giraffidae). A hypothesis of chromosomal evolution involving fission has been outlined by Todd (1975) for the order, but our knowledge of this group is still very incomplete.
Camels. Camels have a disjunct distribution, with four species in South America and two species in the Old World. A study comparing banding patterns in two South American species and the Bactrian camel found the G-banding patterns to be indistinguishable (Bunch, Foote and Maciulis 1985). The distributions of heterochromatin were also indistinguishable, a rather unusual result. The lack of chromosomal divergence despite the geographical isolation is unexpected, and suggests either a very stable karyotype or a relatively short period of isolation, or both.
Cattle family. The karyotypes of the various species of sheep and goats are very similar, with differences attributed to fusions (Bunch, Foote and Spillett 1976). Cattle chromosomes show large homeologies with those of sheep and goats (Schnedl and Czaker 1974). Buckland and Evans (1978), using the goat karyotype as a standard, found nearly complete agreement in banding patterns among several species of bovids, representing three subfamilies. The goat and the horse-like antelope karyotypes were more similar to each other than either was to the cattle karyotype.
A rearrangement which is shared by several similar species is considered to be a good indicator of common ancestry (Rofe 1976). A Y/autosome translocation is found in several species of African cattle-like antelopes (Benirschke et al. 1980), including the eland and the bongo. Differences among the species appear largely due to Robertsonian fusions, with a few tandem fusions and some other unidentified rearrangements. An X/autosome tandem fusion is found in several species of gazelles (Effron et al. 1976, Benirschke et al. 1984). These examples illustrate variability in species which has probably come about relatively recently.
Deer family. One of the most unusual examples of chromosome modification yet discovered is found in the muntjacs, a group of small Asian deer. Two species, the Indian muntjac, Muntiacus muntjak vaginalis, and M. rooseveltorum, share the distinction of having the lowest 2n number known among mammals, six in the female and 7 in the male (Wurster-Hill and Seidel 1985). The Chinese muntjac, M. reevesi, looks very similar but has 2n=46 in both sexes. A comparison of the chromosomes of the Indian and Chinese species (Liming, Yingying and Xingsheng 1980) suggests that essentially the same genetic material is present in each and that the lower number of chromosomes is probably derived by tandem fusion from the 2n=46 karyotype. A fourth species, M. feae, has 2n=13 (Soma et al. 1983). Karyotypes of these species illustrate the usefulness of G-banding in comparing karyotypes and show that caution is in order in drawing phylogenetic conclusions based solely on chromosomal data.
Lagomorphs
Species from both the pika family (Ochotonidae) and the rabbits and hares (family Leporidae) have been studied. The two families appear to share very little or no detectable banding homeologies (Stock 1976), but extensive chromosomal similarities are present among the leporids. Hares (Lepus) have similar karyotypes, while cottontails (Sylvilagus) show considerable variation (Robinson, Elder and Chapman 1983a,b, 1984). The ancestral karyotype for the group appears to be like that of the hares. Other genera of rabbits appear to be related karyotypically to the hares (Robinson and Skinner 1983, Robinson 1980, Stock 1976). Many of the differences can be ascribed to Robertsonian rearrangements.
Rodents
This is the most diverse order of mammals, and a great amount of cytogenetic study has been done with rodents. However, much more remains to be done. Most comparative studies of rodent G-banding have been done with rats and mice, and many families have not yet been studied. A recent bibliography of rodent karyological studies is available (Jotterand-Bellomo 1984).
The G-banding patterns of thirteen species of ground squirrels (Spermophilus) have been published (Nadler et al. 1973, 1975, 1984), and extensive homeologies determined. An interesting geographical pattern has been discovered in this group. The arctic ground squirrel, S. parryi, is found on both sides of the Bering Strait, both populations having identical karyotypes. The arctic ground squirrel populations separate two other species: S. columbianus in North America and S. undulatus in Siberia. The banding patterns of these two species are identical. The highest 2n numbers in the genus are found in the Asian S. xanthoprymnus (2n=42) and the North American S. vigilis (2n=46). These two species differ primarily by two fusions. In contrast to the karyotypic variability of the squirrel genus Spermophilus, most chipmunks (genus Tamias), have very similar karyotypes (Nadler et al. 1977).
By far the largest family of mammals is the mouse family, and a large number of chromosomal studies have been conducted among its members. Only a few studies can be described here. All species of white-footed mice (Peromyscus) studied so far have 48 chromosomes (Robbins and Baker 1981). The most primitive karyotype was proposed to be that of P. boylii. A modified Peromyscus karyotype was proposed to be ancestral for the family by Koop et al. (1984). They noted that karyotypic differences may be more extensive among species in a genus than between genera.
One of the most interesting cases of chromosomal speciation is found in the house mouse (Mus musculus complex). Chromosome numbers range from 22 to 40, with differences due to Robertsonian rearrangements (Gropp and Winking 1981). Such a situation is called a "Robertsonian fan". Several other species from the same subgenus share identical banding patterns with Mus musculus, but at least some species in other subgenera have quite different banding patterns (Hsu, Markvong and Marshall 1978). Another Robertsonian fan has been described in the European mole-vole, Ellobius talpinus (Lyapunova et al. 1984). Here the 2n number varies from 31 to 54 within a geographic distance of only 150 km. All chromosome numbers from 31 to 54 have been found, indicating extensive introgression.
Several genera of native Australian rodents, representing three tribes, have been studied. Each of the three tribes contains a species with a common banding pattern (Baverstock et al. 1983a). These Australian rats have virtually no banding homeologies with species of Rattus, indicating only a distant relationship. One genus (Zyzomys) has apparently had its genome completely rearranged.
Banding patterns of ten genera of murid rodents, mostly of African origin, were compared by Viegas-Pequignot et al. (1983), and an ancestral karyotype proposed for the murid rodents. Several examples were noted in which a particular type of rearrangement appears to have accumulated in a particular lineage (karyotypic orthoselection). This ancestral karyotype for murid rodents was compared with that of a South American cricetid, Akodon arviculoides to test for similarities between the two subfamilies (Viegas-Pequignot et al. 1985). About 40% homeology was claimed. It would be interesting to compare the "ancestral" karyotype proposed by these authors with that given by Koop et al. (1984).
South American hystricomorph rodents. Several families of mostly South American rodents are included in the hystricomorphous rodents. Not many studies of G-band comparisons have been published, and fewer yet in English. A review of unbanded karyotypes of hystricomorphs was published by George and Weir (1974). Three species of Caviidae, representing three genera, were compared by Maia (1984). Differences reported were primarily due to heterochromatin content. Chromosomal speciation appears to be taking place among populations of a superspecies of spiny rats (Proechimys, Family Echimyidae) in Venezuela (Reig et al. 1980). Chromosome numbers range from 42 to 62. Differences are due to Robertsonian fusions, except for the extreme chromosome numbers, where pericentric inversions are also involved.
Miscellaneous rodent families. Five species of gundis (family Ctenodactylidae), representing four genera, have been shown to have similar chromosomal banding patterns (George 1979a). Differences can be explained by a pericentric inversion, and perhaps several very small translocations. The same author (George 1979b) found very close similarity in the banding patterns of two species of African mole-rats (family Bathyergidae). This contrasts sharply with the variability seen in some other families of burrowing rodents and casts doubt on the hypothesis that chromosomal evolution is especially promoted by the kind of social structure found in burrowing rodents (Wilson et al. 1975; see also Gileva 1983).
A CREATIONIST VIEWPOINT
A challenge for creationism
Although much remains to be learned about the meaning of chromosomal structure, enough data on chromosomal comparisons have been gathered to raise some important questions for creationists. That changes have occurred in organisms since creation is not in question, but the extent of those changes is uncertain.
Species which are similar morphologically generally have similar karyotypes, although there are significant exceptions (e.g., see Liming, Yingying and Xingsheng 1980). This is quite reasonable if species with similar morphology (e.g., in the same genus) are thought of as being related through common ancestry. The occasional exception merely shows that chromosomes can be extensively rearranged with no significant morphological effect.
More problematic is the finding that species which are quite different morphologically may have similar karyotypes. The chromosomal similarities among many of the Australian marsupials (Rofe and Hayman 1985), between goats and giraffes (Buckland and Evans 1978), between seals and terrestrial carnivores (Arnason 1977) and between humans and the great apes (Yunis and Prakesh 1982) raise some significant questions for creationists. Perhaps the two most important questions are:
- To what extent has morphological change occurred in mammals, and by what mechanisms? and
- What is the relationship between humans and apes?
These questions will be amplified below, and then various hypotheses regarding the relationship of chromosomal evidence to these questions will be discussed.
Problem 1. The extent and mechanism of morphological change. There is circumstantial evidence that mammal species may change significantly in their morphology. This evidence comes from the study of island populations (e.g., Lawlor 1982, Simpson 1956), from the results of selective breeding of domestic animals (e.g., Wayne 1986), from the ability of some animals to hybridize (Van Gelder 1977), and from distributional patterns of living mammals (Darwin 1859). However, there seem to be limits on the amount of morphological change possible (Lester and Bohlin 1984).
If groups such as the Australian marsupial families are considered to share a common ancestry in spite of their diverse morphology, one is challenged to propose some mechanism by which such change could be brought about. The standard neodarwinian gradualistic explanation for morphological change is that small changes arise by mutation and accumulate over time by natural selection to produce large changes (e.g., see Charlesworth et al. 1982). However, the lack of fossil intermediates, or even conceivable intermediate stages, has led many scientists to search for other explanations. Several alternative mechanisms for macroevolutionary changes have been proposed (e.g., Gould 1977; Oster and Alberch 1982; Wilson, Maxson and Sarich 1974; Wright 1982), but none has been satisfactory. The possible role of chromosomal rearrangements in speciation was discussed in a previous article (Gibson 1984).
For creationists, the origin of diversity in mammals is an important question. If enough morphological change has occurred since the Genesis flood to explain the origin of diversity among marsupials, it seems reasonable to think that the same amount of change could also have happened among placental mammals, although placentals as a group do not have such similar chromosomal banding patterns as marsupials. However, in the absence of a plausible genetic mechanism for creating new adaptations, creationists are somewhat skeptical that such changes have occurred, even though there is no scriptural prohibition against large changes in species.
If the marsupials are considered to be unrelated, then one has the problem of explaining why they share so many unique characteristics, including chromosomal similarities and such structural traits as their reproductive anatomy, the presence of epipubic bones, and the inflection of the angular process of the lower jaw. Their geographic distribution is also difficult to explain.
Problem 2. The relationship of man and the apes. Questions concerning the origin and nature of man have deep philosophical significance. Evolutionists have long held that humans and apes share a common ancestry, a belief based largely on morphological similarities. Fossil discoveries have not clarified the picture, but seemingly have made it more confused, perhaps due to the subjective nature of interpreting the fossils (Washburn 1973). However, striking similarities have been discovered between apes and humans in their proteins (Bruce and Ayala 1979), their chromosomes (Yunis and Prakesh 1982), and in their DNA (Sibley and Ahlquist 1984).
To say that humans and apes are not related by common descent is to emphasize their difference in anatomy and behavior, and to downgrade the importance of their similarities in anatomy, biochemistry and chromosomes. Although the human karyotype is considered to be closest to the ancestral condition for humans and apes (Yunis and Prakesh 1982), I am not aware of any serious examination of the possibility that humans might be ancestral to apes.
The meaning of chromosomal similarity
As an explanation for similarities in chromosomal structure, four distinct possibilities come to mind, each presented as a separate hypothesis below.
Hypothesis 1. Chromosomal similarities are the result of common design. This would mean that organisms which are similar morphologically were created with similar karyotypes, just as they were created with similar anatomical and biochemical features. If the karyotypes have not undergone much change since creation, we should be able to see the similarities. Whether a karyotype should be shared by all mammals or only by those with some degree of morphological similarities is uncertain.
If a karyotype is shared only by species with similar morphology, one might infer that the structure of the chromosomes is somehow related to the morphology of the organism. It is true that, in general, groups of species with similar G-banding patterns are also similar morphologically. However, it is known that major changes in the karyotype, as shown by G-banding, do not cause morphological change (e.g., see Baker, Bickham and Arnold 1985). It is also known that different types of chromosomal change may be found in groups of species which could plausibly have a common ancestry (e.g., see Koop et al. 1984). These facts cast doubt on any fixed relationship between chromosome morphology and anatomical morphology, although they do not disprove a possible original relationship between them.
There have been some suggestions that karyotypic structure has adaptive significance (Baker et al. 1983, Kiel et al. 1985), but this has not been demonstrated conclusively. It is of interest to note that there is frequently a correlation between anatomical distinctiveness and karyotypic distinctiveness between groups at high taxonomic levels.
Hypothesis 2. Chromosomal similarities are exclusively the result of common ancestry. According to this hypothesis, if two species have similar chromosomes (including banding patterns), they are related. This would require that each original species group was created with its own unique karyotype. If this hypothesis is correct, one must accept a common ancestry for apes and humans, for at least the majority of the marsupials, and for cattle, goats, antelope and giraffes. Acceptance of this hypothesis is the basis for phylogenies based on chromosomal similarities.
There are problems with this hypothesis. One is that despite some circumstantial evidence for major morphological change in mammals, no mechanism is known which would account for the kind of changes here suggested. Among the Australian marsupials, for example, there are considerable morphological differences between the wombat, the bandicoot and the "native cat". A second problem is that in comparing banding patterns which are similar but may not be identical, preconceived ideas of ancestry can bias one's conclusions (e.g., see above under Primates). If one assumes that two similar species do in fact have a common ancestor, then one is committed to finding a way of matching the banding patterns. In view of the subjectivity involved in matching chromosomal banding patterns, one might wonder about the significance of a 25% or 50% match of banding patterns, especially it no entire arms can be matched.
Hypothesis 3. Chromosomal similarities are due to random changes which happen to produce the same banding pattern in different species. This hypothesis implies that each originally created species had a unique karyotype. It also implies that chromosomal similarities have no real significance. If a series of patterns is made by randomly arranging dark and light bands, it is inevitable that some patterns will be repeated by chance. Thus the similarities of chromosome banding in humans and apes and within some other groups could be held to be merely a result of chance.
This hypothesis does not seem reasonable for two reasons. Similar chromosome banding patterns are not found randomly distributed throughout all taxonomic groups, but rather are found in groups which share morphological similarities. This argues strongly against any random cause of the banding patterns. In fact, it is known that breakage points in chromosomes are not random (Jacky, Beek and Sutherland 1983), and so changes in a karyotype will not be random. The phenomenon of "karyotypic orthoselection" also shows that chromosomal changes are non-random. If chromosomes have non-random breakage points, similar (parallel) rearrangements could occur independently in similar karyotypes (see below), but it is unlikely that convergent events would occur in different karyotypes to produce similar results.
Hypothesis 4. Chromosomal similarities are the result of non-random changes due to viruses or transposable elements. This hypothesis requires that either 1) karyotypes which were once different have been caused to become similar, or 2) karyotypes which were once similar have been changed in a similar way, due to the action of transposable elements or some similar mechanism.
Transposable elements (TEs) are known to increase the rate of chromosomal rearrangements in a non-random way (Nevers and Saedler 1977). But if the karyotypes were substantially different, there is no reason to expect them to change to be the same, since the insertion sites of TEs appear to be at least partially sequence-dependent (see Inouye, Yuki and Saigo 1984; Shapiro 1979).
Similar (parallel) changes do sometimes occur in similar species, as has been shown in several studies (e.g., Robbins and Baker 1981; Baker, Koop and Haiduk 1983; Baker, Bickham and Arnold 1985; Searle 1984). A common ancestry is plausible in each of these cases, and it is likely that the rather similar species have undergone numerous chromosomal changes during speciation, some of which happened to be the same.
It seems more likely that transposable elements could cause karyotypes which were originally similar to become different. This could occur if different species were infected by different TEs or retroviruses having different effects on the genome (see Rose and Doolittle 1983). It seems possible that different TEs might affect a genome in different ways. As a hypothetical example, it seems possible that the ancestors of oryzomine and peromyscine rodents (groups of rats and mice) could originally have had similar karyotypes when infected by different TEs. The TE(s) infecting the peromyscine lineage might have caused a series of heterochromatin additions and pericentric inversions, while the TE infecting the oryzomine lineage might have caused a series of fusions.
CONCLUSIONS
It is possible that chromosomal similarities have different explanations in different groups of animals. It this is true, then one must be cautious in using chromosomal comparisons to determine relationships. Nevertheless, chromosomal data can serve as a useful check on data from other sources.
Hypothesis 3, that chromosomal similarities are due to random chromosomal rearrangements which happen to produce similar banding patterns, is not reasonable, for reasons discussed above. Hypothesis 2, that chromosomal similarities are exclusively the result of common ancestry, does not seem consistent with creation theory and does not seem a necessary conclusion from the scientific data. The fact that very large genomic rearrangement does not seem to affect morphology, and yet animals with different body plans ("Bauplan") appear to have very different kinds of karyotypes suggests to this writer that some different groups had different starting points and do not share a common ancestry.
Hypotheses 1 and 4 seem consistent with both creation theory and the evidence available. It seems likely that species which were morphologically similar were created with similar chromosomes, reflecting their genetic similarity. It is evident that large changes have occurred in chromosomes since creation. These changes have often resulted in karyotypic divergence and have contributed to the multiplication of species.
Chromosomal rearrangements seem to occur so frequently that one would expect to find very little banding homeology between species which supposedly diverged long ages ago, such as the marsupials. The existence of numerous banding homeologies can be explained as the result of a common design which has been preserved only because a relatively short time has been available for changes to occur.
How much anatomical change has occurred since creation is still an unanswered question. Chromosomal comparisons suggest that new genera may have arisen since creation, for example among the antelopes which share a Y/autosome translocation (Benirschke et al. 1980). Whether larger changes have occurred cannot be determined from chromosomal studies. At the present time there is no known mechanism by which changes in organisms can take place which are large enough to account for the differences among, for example, the Australian marsupials or the various families of artiodactyls (cattle, giraffes, deer). The absence of fossil evidence linking different groups by a common ancestry, together with the lack of biological evidence of a mechanism for such change, seem consistent with the hypothesis that they have separate ancestries.
LITERATURE CITED
- Arnason, U. 1974. Comparative chromosome studies in Cetacea. Hereditas 77:1-36.
- Arnason, U. 1977. The relationship between the four principal pinniped karyotypes. Hereditas 87:227-242.
- Arnason, U., K. Benirschke, J. G. Mead, and W. W. Nichols. 1977. Banded karyotypes of three whales: Mesoplodon europaeus, M. carlhubbsi and Balaenoptera acutorostrata. Hereditas 87:189-200.
- Atchley, W. R. 1972. The chromosome karyotype in estimation of lineage relationships. Systematic Zoology 21:199-209.
- Baker, R. J. and J. W. Bickham. 1980. Karyotypic evolution in bats: evidence of extensive and conservative chromosomal evolution in closely related taxa. Systematic Zoology 29:239-253.
- Baker, R. J., R. K. Barnett, and I. F. Greenbaum. 1979. Chromosomal evolution in grasshopper mice (Onychomys: Cricetidae). Journal of Mammalogy 60:297-306.
- Baker, R. J., J. W. Bickham, and M. L. Arnold. 1985. Chromosomal evolution in Rhogeesa (Chiroptera: Vespertilionidae): Possible speciation by centric fusions. Evolution 39:233-243.
- Baker, R. J., B. F. Koop, and M. W. Haiduk. 1983. Resolving systematic relationships with G-bands: a study of five genera of South American cricetine rodents. Systematic Zoology 32:403-416.
- Baker, R. J., R. K. Chesser, B. F. Koop, and R. A. Hoyt. 1983. Adaptive nature of chromosomal rearrangement: differential fitness in pocket gophers. Genetica 61:161-164.
- Baverstock, P. R., C. H. S. Watts, M. Gelder, and A. Jahnke. 1983a. G-banding homologies of some Australian rodents. Genetica 60:105-117.
- Baverstock, R. P., M. Adams, M. Archer, N. L. McKenzie, and R. How. 1983b. An electrophoretic and chromosomal study of the dasyurid marsupial genus Ningaui Archer. Australian Journal of Zoology 31:381-392.
- Benirschke, K., D. Ruedi, H. Muller, A. T. Kumamoto, K. L. Wagner, and H. S. Downes. 1980. The unusual karyotype of the lesser kudu, Tragelaphus imberbis. Cytogenetics and Cell Genetics 26:85-92.
- Benirschke, K., A. T. Kumamoto, J. H. Olsen, M. M. Williams, and J. Oosterhuis. 1984. On the chromosomes of Gazella soemmeringi Cretzschmar, 1826. Zeitschrift für Saugetierkunde 49:368-373.
- Benton, M. J. 1985. First marsupial fossil from Asia. Nature 318:313.
- Bick, Y. A. E. and B. G. Sharman. 1975. The chromosomes of the platypus (Ornithorhynchus: Monotremata). Cytobios 14:17-28.
- Bickham, J. W. 1979a. Banded karyotpyes of 11 species of American bats (genus Myotis). Cytologia 44:789-797.
- Bickham, J. W. 1979b. Chromosomal variation and evolutionary relationships of vespertilionid bats. Journal of Mammalogy 60:350-363.
- Bruce, E. J. and F. J. Ayala. 1979. Phylogenetic relationship between man and the apes: electrophoretic evidence. Evolution 33:1040-1056.
- Buckland, R. A. and H. J. Evans. 1978. Cytogenetic aspects of phylogeny in the Bovidae. I. G-banding. Cytogenetics and Cell Genetics 21:42-63.
- Bunch, T. D., W. C. Foote, and J. J. Spillett. 1976. Translocations of acrocentric chromosomes and their implications in the evolution of sheep (Ovis). Cytogenetics and Cell Genetics 17:122-136.
- Bunch, T. D., W. C. Foote, and A. Maciulis. 1985. Chromosome banding pattern homeologies and NORs for the Bactrian camel, guanaco, and llama. The Journal of Heredity 76:115-118.
- Bush, G. L. 1981. Stasipatric speciation and rapid evolution in animals. In W. R. Atchley and D. S. Woodruff, eds. Evolution and Speciation: Essays in Honor of M. J. D. White, pp. 201-218. Cambridge University Press, Cambridge.
- Casartelli, C., S. R. Rogatto, and I. Ferrari. 1986. Cytogenetic analysis of some Brazilian marsupials (Didelphidae: Marsupialia). Canadian Journal of Genetics and Cytology 28:21-29.
- Charlesworth, B., R. Lande, and M. Slatkin. 1982. A neo-Darwinian commentary on macroevolution. Evolution 36:474-498.
- Couturier, J., E. Razafimahatratra, B. Dutrillaux, S. Warter, and Y. Rumpler. 1986. Chromosomal evolution in the Malagasy Carnivora. I. R-banding studies of Cryptoprocta ferox, Fossa fossana, Galidia elegans, and Mungotictis decemlineata. Cytogenetics and Cell Genetics 41:1-8.
- Darwin, C. R. 1869. The origin of species. John Murray, London. Undated reprint from the 6th London edition. National Library Association of Chicago.
- Duffy, P. A. 1972. Chromosome variation in Peromyscus: a new mechanism. Science 176:1333-1334.
- Dutrillaux, B. 1979. Chromosomal evolution in Primates: tentative phylogeny from Microcebus murinus (prosimian) to man. Human Genetics 48:251-314.
- Dutrillaux, B., E. Viegas-Pequignot, J. Couturier, and G. Chauvier. 1978. Identity of euchromatic bands from man to Cercopithecidae. Human Genetics 45:283-296.
- Dutrillaux, B. and J. Couturier. 1981. The ancestral karyotype of platyrrhine monkeys. Cytogenetics and Cell Genetics 30:232-242.
- Dutrillaux, B. and J. Couturier. 1983. The ancestral karyotype of Carnivora: comparison with that of platyrrhine monkeys. Cytogenetics and Cell Genetics 35:200-208.
- Dutrillaux, B., J. Couturier and A.-M. Fosse. 1980. The use of high resolution banding in comparative cytogenetics: comparison between man and Lagothrix lagotricha (Cebidae). Cytogenetics and Cell Genetics 27:45-51.
- Dutrillaux, B., J. Couturier, S. Warter, and Y. Rumpler. 1982. Chromosomal evolution in 'Lemurs'. VI. Chromosomal banding studies of Galago senegalensis, Galago alleni, Galago demidovii and Euoticus elegantulus. Folia primatologica 37:280-296.
- Effron, M., M. H. Bogart, A. T. Kumamoto, and K. Benirschke. 1976. Chromosome studies in the mammalian subfamily Antilopinae. Genetica 46:419-444.
- Elder, F. F. B. 1980. Tandem fusion, centric fusion, and chromosomal evolution in the cotton rats, genus Sigmodon. Cytogenetics and Cell Genetics 26:199-210.
- Engstrom, M. D. and J. W. Bickham. 1982. Chromosome banding and phylogeny of the golden mouse Ochrotomys nuttalli. Genetica 59:119-126.
- Estop, A. M., J. J. Garver, and P. L. Pearson. 1978. Further studies on the comparative karyology of the African green and rhesus monkeys. Genetica 49:131-138.
- Farris, J. S. 1978. Inferring phylogenetic trees from chromosome inversion data. Systematic Zoology 27:275-284.
- Fredga, K. 1972. Comparative chromosome studies in mongooses (Carnivora, Viverridae). I. Idiograms of 12 species and karyotype evolution in Herpestinae. Hereditas 71:1-74.
- Galbreath, G. J. 1983. Karyotypic evolution in Aotus. American Journal of Primatology 4:245-251.
- George, W. 1979a. Conservatism in the karyotypes of two African mole rats (Rodentia, Bathyergidae). Zeitschrift für Saugetierkunde 44:278-285.
- George, W. 1979b. The chromosomes of the hystricomorphous family Ctenodactylidae (Rodentia: ?Sciuromorpha) and their bearing on the relationships of the four living genera. Zoological Journal of the Linnean Society 65:261-280.
- George, W. and B. J. Weir. 1974. Hystricomorph chromosomes. Symposia of the Zoological Society of London 34:79-108.
- Gibson, L. J. 1984. Chromosomal changes in mammalian speciation: a literature review. Origins. 11:67-89.
- Gileva, E. A. 1983. A contrasted pattern of chromosome evolution in two genera of lemmings, Lemmus and Dicrostonyx (Mammalia, Rodentia). Genetica 60:173-179.
- Gould, S. J. 1977. The return of hopeful monsters. Natural History 76:22-30.
- Gropp, A. and H. Winking. 1981. Robertsonian translocations: cytology, meiosis, segregation patterns and biological consequences of heterozygosity. Symposia of the Zoological Society of London 47:141-181.
- Haiduk, M. W. and R. J. Baker. 1982. Cladistical analysis of G-banded chromosomes of nectar-feeding bats (Glossophaginae: Phyllostomatidae). Systematic Zoology 31:252-265.
- Haiduk, M. W., R. U. Baker, L. W. Robbins, and D. A. Schlitter. 1981. Chromosomal evolution in African megachiroptera: G- and C-band assessment of the magnitude of change in similar standard karyotypes. Cytogenetics and Cell Genetics 29:221-232.
- Hayman, D. L. 1977. Chromosome number-constancy and variation. In B. Stonehouse and D. Gilmore, eds. The Biology of Marsupials, pp. 27-48. University Park Press, Baltimore.
- Hinesley, L. L. 1979. Systematics and distribution of two chromosome forms in the southern grasshopper mouse, genus Onychomys. Journal of Mammalogy 60:117-128.
- Hsu, T. C., A. Markvong, and J. T. Marshall. 1978. G-band patterns of six species of mice belonging to the subgenus Mus. Cytogenetics and Cell Genetics 20:304-307.
- Inouye, S., S. Yuki, and K. Saigo. 1984. Sequence-specific insertion of the Drosophila transposable genetic element 17.6. Nature 310:332-333.
- Jacky, P. B., B. Beek, and G. R. Sutherland. 1983. Fragile sites in chromosomes: possible model for the study of spontaneous chromosome breakage. Science 220:69-70.
- Jotterand-Bellomo, M. 1984. New developments in vertebrate cytotaxonomy VII. Les chromosomes des Rongeurs (ordre Rodentia Bowdich, 1821). Genetica 64:3-64.
- Kiel, K. von, H. Hameister, I. E. Somssich, and S. Adolph. 1985. Early replication banding reveals a strongly conserved functional pattern in mammalian chromosomes. Chromosoma 93:69-76.
- Koop, B. F., R. J. Baker, M. W. Haiduk, and M. D. Engstrom. 1984. Cladistical analysis of primitive G-band sequences for the karyotype of the ancestor of the Cricetidae complex of rodents. Genetica 64:199-208.
- Lalley, P. A. and V. A. McKusick. 1985. Report of the committee on comparative mapping. Cytogenetics and Cell Genetics 40:536-566.
- Lawlor, T. E. 1982. The evolution of body size in mammals: evidence from insular populations in Mexico. American Naturalist 119:54-72.
- Lee, M. R. and L. K. Martin. 1980. Mastomys (=Praomys) natalensis is not a Rattus (Mammalia: Rodentia): karyological evidence. Cytogenetics and Cell Genetics 28:95-103.
- Lester, L. P. and R. G. Bohlin. 1984. The natural limits to biological change. Zondervan, Grand Rapids, Michigan.
- Liming, S., Y. Yingying, and D. Xingsheng. 1980. Comparative cytogenetic studies on the red muntjac, Chinese muntjac, and their F1 hybrids. Cytogenetics and Cell Genetics 26:22-27.
- Lyapunova, E. A., S. B. Ivnitskii, V. P. Korablev, and I. Yu. Yanina. 1984. A complete Robertsonian complex of the chromosomal forms in the mole-vole superspecies Ellobius talpinus. Doklady Biological Sciences 274:87-90. Translated from Doklady Akademii Nauk SSSR 274(5):1209-1213, February, 1984.
- Ma, N. S. F. 1981. Chromosome evolution in the owl monkey, Aotus. American Journal of Physical Anthropology 54:293-303.
- Maia, V. 1984. Karyotypes of three species of Caviinae (Rodentia, Caviidae). Experientia 40:564-566.
- Mascarello, J. T., A. D. Stock, and S. Pathak. 1974. Conservation in the arrangement of genetic material in rodents. Journal of Mammalogy 55:695-704.
- Matthey, R. 1973a. The chromosome formulae of eutherian mammals. In A. B. Chiarelli and E. Capanna, eds. Cytotaxonomy and Vertebrate Evolution, pp. 531-616. Academic Press, New York.
- Matthey, R. 1973b. Les nombres diploides des eutheriens. Mammalia 37:394-421.
- Mudry de Pargament, M. D., S. Brieux de Salum, and O. J. Colillas. 1984. Bibliography of cytogenetic studies of Platyrrhini. Journal of Human Evolution 13:217-221.
- Murtagh, C. E. 1977. A unique cytogenetic system in monotremes. Chromosoma 65:37-57.
- Nadler, C. F. and R. S. Hoffmann. 1974. Chromosomes of the African ground squirrel, Xerus rutilus (Rodentia: Sciuridae). Experientia 30:889-891.
- Nadler, C. F., L. W. Turner, R. S. Hoffmann, and L. Deutsch. 1973. Chromosomes and Giemsa-bands of the Idaho spotted ground squirrel, Spermophilus brunneus (Howell). Experientia 29:893-894.
- Nadler, C. F., E. A. Lyapunova, R. S. Hoffmann, N. N. Vorontsov, and N. A. Malygina. 1975. Chromosomal evolution in Holarctic ground squirrels (Spermophilus). I. Giemsa-band homologies in Spermophilus columbianus and S. undulatus. Zeitschrift für Saugetierkunde 40:1-7.
- Nadler, C. F., R. S. Hoffmann, J. H. Honacki, and D. Pozin. 1977. Chromosomal evolution in chipmunks, with special emphasis on A and B karyotypes of the subgenus Neotamias. American Midland Naturalist 98:343-353.
- Nadler, C. F., E. A. Lyapunova, R. S. Hoffmann, N. N. Vorontsov, L. L. Shaitarova, and Y. M. Borisov. 1984. Chromosomal evolution in Holarctic ground squirrels (Spermophilus). II. Giemsa-band homologies of chromosomes and the tempo of evolution. Zeitschrift für Saugetierkunde 49:78-90.
- Nash, W. G. and S. J. O'Brien. 1982. Conserved regions of homologous G-banded chromosomes between orders in mammalian evolution: carnivores and primates. Proceedings of the National Academy of Sciences USA 79:6631-6635.
- Nevers, P. and H. Saedler. 1977. Transposable genetic elements as agents of gene instability and chromosomal rearrangements. Nature 268:109-115.
- Nowak, R. M. and J. L. Paradiso. 1983. Walker's mammals of the world. 2 vols. Johns Hopkins, Baltimore.
- O'Brien, S. J. and W. G. Nash. 1982. Genetic mapping in mammals: chromosome map of domestic cat. Science 216:257-265.
- O'Brien, S. J., W. G. Nash, D. E. Wildt, M. E. Bush, and R. E. Benveniste. 1985. A molecular solution to the riddle of the giant panda's phylogeny. Nature 317:140-144.
- Olert, J. and M. Schmid. 1978. Comparative analysis of karyotypes in European shrew species. I. The sibling species Sorex araneus and S. gemellus: Q-bands, G-bands, and position of NORs. Cytogenetics and Cell Genetics 20:308-322.
- Pathak, S. and A. D. Stock. 1976. Giemsa-banding and the identification of the Y/autosome translocation in the African marsh mongoose, Atilax paludinosus (Carnivora, Viverridae). Cytogenetics and Cell Genetics 16:487-494.
- Patton, J. C. and R. J. Baker. 1978. Chromosomal homology and evolution of phyllostomatoid bats. Systematic Zoology 27:449-462.
- Petit, D., J. Couturier, E. Viegas-Pequignot, M. Lombard, and B. Dutrillaux. 1984. Tres grande similitude entre le caryotype ancestral des ecureuils (Rongeurs) et celui des Primates et de Carnivores. Annales de Genetique 27:201-212.
- Ponsa, M., R. Miro, A. M. Estop, and J. Egozcue. 1981. Banding patterns of the chromosomes of Erythrocebus patas (Schreber 1774) compared to other primate species. Genetica 56:39-45.
- Reig, O. A., M. Aguilera, M. A. Barros, and M. Useche. 1980. Chromosomal speciation in a rassenkreis of Venezuelan spiny rats (Genus Proechimys, Rodentia, Echimyidae). Genetica 52/53:291-312.
- Robbins, L. W. and R. J. Baker. 1981. An assessment of the nature of chromosomal rearrangements in 18 species of Peromyscus (Rodentia: Cricetidae). Cytogenetics and Cell Genetics 31:194-202.
- Robinson, T. J. 1980. Comparative chromosome studies in the family Leporidae (Lagomorpha, Mammalia). Cytogenetics and Cell Genetics 28:64-70.
- Robinson, T. J. and J. D. Skinner. 1983. Karyology of the riverine rabbit, Bunolagus monticularis, and its taxonomic implications. Journal of Mammalogy 64:678-681.
- Robinson, T. J., F. F. B. Elder, and J. A. Chapman. 1983a. Karyotypic conservatism in the genus Lepus (order Lagomorpha). Canadian Journal of Genetics and Cytology 25:540-544.
- Robinson, T. J., F. F. B. Elder, and J. A. Chapman. 1983b. Evolution of chromosomal variation in cottontails, genus Sylvilagus (Mammalia: Lagomorpha): S. aquaticus, S. floridanus, and S. transitionalis. Cytogenetics and Cell Genetics 35:216-222.
- Robinson, T. J., F. F. B. Elder, and J. A. Chapman. 1984. Evolution of chromosomal variation in cottontails, genus Sylvilagus (Mammalia: Lagomorpha). Cytogenetics and Cell Genetics 38:282-289.
- Rofe, R. H. 1976. G-banded chromosomes and the evolution of Macropodidae. Australian Mammalogy 2:53-63.
- Rofe, R. and D. Hayman. 1985. G-banding evidence for a conserved complement in Marsupialia. Cytogenetics and Cell Genetics 39:40-50.
- Rose, M. R. and W. F. Doolittle. 1983. Molecular biological mechanisms of speciation. Science 220:157-162.
- Rumpler, Y. and B. Dutrillaux. 1976. Chromosomal evolution in Malagasy lemurs. I. Chromosome banding studies in the genuses Lemur and Microcebus. Cytogenetics and Cell Genetics 17:268-281.
- Rumpler, Y. and B. Dutrillaux. 1978. Chromosomal evolution in Malagasy lemurs. III. Chromosome banding studies in the genus Hapalemur and the species Lemur catta. Cytogenetics and Cell Genetics 21:201-211.
- Rumpler, Y. and B. Dutrillaux. 1979. Chromosomal evolution in Malagasy lemurs. IV. Chromosome banding studies in the genuses Phaner, Varecia, Lemur, Microcebus, and Cheirogalus. Cytogenetics and Cell Genetics 24:224-232.
- Rumpler, Y., J. Couturier, S. Warter, and B. Dutrillaux. 1983a. The karyotype of Galago crassicaudatus is ancestral for Lorisiforms. Folia primatologica 40:227-231.
- Rumpler, Y., J. Couturier, S. Warter, and B. Dutrillaux. 1983b. Chromosomal evolution in Malagasy lemurs. VII. Phylogenic relationships between Propithecus, Avahi (Indridae), Microcebus (Cheirogaleidae), and Lemur (Lemuridae). Cytogenetics and Cell Genetics 36:542-546.
- Rumpler, Y., B. Ishak, S. Warter, and B. Dutrillaux. 1985. Chromosomal evolution in Malagasy lemurs. VIII. Chromosome banding studies of Lepilemur ruficaudatus, L. leucopus, and L. septentrionalis. Cytogenetics and Cell Genetics 39:194-199.
- Ryder, O. A., N. C. Epel, and K. Benirschke. 1978. Chromosome banding studies of the Equidae. Cytogenetics and Cell Genetics 20:323-350.
- Schnedl, W. and R. Czaker. 1974. Centromeric heterochromatin and comparison of G-banding in cattle, goat, and sheep chromosomes (Bovidae). Cytogenetics and Cell Genetics 13:246-255.
- Seabright, M. 1972. The use of proteolytic enzymes for the mapping of structural rearrangements in the chromosomes of man. Chromosoma 36:204-210.
- Searle, J. B. 1984. Hybridization between Robertsonian karyotypic races of the common shrew Sorex araneus. Experientia 40:876-878.
- Shapiro, J. A. 1979. Molecular model for the transposition and replication of bacteriophage Mu and other transposable elements. Proceedings of the National Academy of Sciences USA 76:1933-1937.
- Sharman, G. B. 1973. The chromosomes of non-Eutherian mammals. In A. B. Chiarelli and E. Capanna, eds. Cytotaxonomy and Vertebrate Evolution, pp. 485-530. Academic Press, New York.
- Sharman, G. B. 1982. Karyotypic similarities between Dromiciops australis (Micro-biotheriidae, Marsupialia) and some Australian marsupials. In M. Archer, ed. Carnivorous Marsupials, pp. 711-714. Royal Zoological Society of New South Wales, Sydney.
- Sibley, C. G. and J. E. Ahlquist. 1984. The phylogeny of the hominoid primates, as indicated by DNA-DNA hybridization. Journal of Molecular Evolution 20:2-15.
- Simpson, G. G. 1956. Zoogeography of West Indian land mammals. American Museum of Natural History Novitates 1759:1-28.
- Soma, H., H. Kada, K. Mtayoshi, Y. Suzuki, C. Meckvichal, A. Mahannop, and B. Vatanaromya. 1983. The chromosomes of Muntiacus feae. Cytogenetics and Cell Genetics 35:156-158.
- Spotorno, A. E. 1977. Phylogenetic partitioning of banded karyotypes in mammals: a model of cladistic analysis. Third Latin American Congress of Genetics (UNESCO), pp. 179-187. Montevideo.
- Stock, A. D. 1976. Chromosome banding pattern relationships of hares, rabbits, and pikas (order Lagomorpha). A phyletic interpretation. Cytogenetics and Cell Genetics 17:78-88.
- Stock, A. D. and T. C. Hsu. 1973. Evolutionary conservation in arrangement of genetic material. Chromosoma 43:211-224.
- Sun, N. C., C. R. Y. Sun, and T. Ho. 1978a. Chimpanzee chromosome 12 is homologous to human chromosome arm 2q. Cytogenetics and Cell Genetics 22:594-597.
- Sun, N. C., C. R. Y. Sun, and T. Ho. 1978b. Chimpanzee chromosome 13 is homologous to human chromosome arm 2p. Cytogenetics and Cell Genetics 22:598-601.
- Todd, N. B. 1975. Chromosomal mechanisms in the evolution of artiodactyls. Paleobiology 1:175-188.
- Van Gelder, R. G. 1977. Mammalian hybrids and generic limits. American Museum of Natural History Novitates 2635:1-25.
- Viegas-Pequignot, E., B. Dutrillaux, M. Prod'Homme, and F. Petter. 1983. Chromosomal phylogeny of Muridae: a study of 10 genera. Cytogenetics and Cell Genetics 35:269-278.
- Viegas-Pequignot, E., S. Kasahara, Y. Yassuda, and B. Dutrillaux. 1985. Major chromosome homeologies between Muridae and Cricetidae. Cytogenetics and Cell Genetics 39:258-261.
- Washburn, S. L. 1973. The evolution game. Journal of Human Evolution 2:557-561.
- Wayne, R. K. 1986. Cranial morphology of domestic and wild canids: the influence of development on morphological change. Evolution 40:243-261.
- White, M. J. D. 1973. Animal cytology and evolution. Cambridge University Press, London.
- White, M. J. D. 1978. Modes of speciation. W. H. Freeman, San Francisco, California.
- Wilson, A. C., L. R. Maxson, and V. M. Sarich. 1974. Two types of molecular evolution. Evidence from studies of interspecific hybridization. Proceedings of the National Academy of Sciences USA 71:2843-2847.
- Wilson, A. C., V. M. Sarich, and L. R. Maxson. 1974. The importance of gene rearrangement in evolution: evidence from studies on rates of chromosomal, protein, and anatomical evolution. Proceedings of the National Academy of Sciences USA 71:3028-3030.
- Wilson, A. C., G. L. Bush, S. M. Case, and M.-C. King. 1975. Social structuring of mammalian populations and rate of chromosomal evolution. Proceedings of the National Academy of Sciences USA 72:5061-6065.
- Wright, S. 1982. The shifting balance theory and macroevolution. Annual Review of Genetics 16:1-19.
- Wurster-Hill, D. H. and M. Bush. 1980. The interrelationship of chromosome banding patterns in the giant panda (Ailuropoda melanoleuca), hybrid bear (Ursus middendorfi ´ Thalarctos maritimus), and other carnivores. Cytogenetics and Cell Genetics 27:147-154.
- Wurster-Hill, D. H. and C. W. Gray. 1973. Giemsa banding patterns in the chromosomes of twelve species of cats. Cytogenetics and Cell Genetics 12:377-397.
- Wurster-Hill, D. H. and C. W. Gray. 1975. The interrelationships of chromosome banding patterns in procyonids, viverrids, and felids. Cytogenetics and Cell Genetics 15:306-331.
- Wurster-Hill, D. H. and B. Seidel. 1985. The G-banded chromosomes of Roosevelt's muntjac, Muntiacus rooseveltorum. Cytogenetics and Cell Genetics 39:75-76.
- Yates, T. L., R. J. Baker, and R. K. Barnett. 1979. Phylogenetic analysis of karyological variation in three genera of peromyscine rodents. Systematic Zoology 28:40-48.
- Yonenaga-Yassuda, Y., S. Kasahara, M. J. Souze, and M. L'Abbate. 1982. Constitutive heterochromatin, G-bands and nucleolus-organizer regions in four species of Didelphidae (Marsupialia). Genetica 58:71-77.
- Young, G. J., J. A. M. Graves, I. Barbieri, P. A. Woolley, D. W. Cooper, and M. Westerman. 1982. The chromosomes of dasyurids (Marsupialia). In M. Archer, ed. Carnivorous Marsupials, pp. 783-795. Royal Zoological Society of New South Wales, Sydney.
- Yunis, J. J. and O. Prakesh. 1982. The origin of man: a chromosomal pictorial legacy. Science 215:1525-1530.
- Zima, J. 1982. Chromosomal homology in the complements of bats of the family Vespertilioniidae: 2. G-band karyotypes of some Myotis, Eptesicus and Pipistrellus species. Folia zoologica 31:31-36.
COVER PICTURES

Front Cover: The chimpanzee (Pan troglodytes). The banding patterns of chimpanzee chromosomes show extensive matching with those of human chromosomes. For a discussion of the significance of chromosomal matching, see the article by James Gibson, pp. 9-35.
Back Cover: The markhor (Capra falconeri), a species of goat native to the mountains of Afghanistan, Pakistan and nearby regions. G-banding patterns of goat chromosomes match well with those of many other species of artiodactyls (mammals with split hoofs), leading to the suggestion that many artiodactyls descended from an ancestor with a goat-like karyotype. Cover photographs taken at the Los Angeles Zoo by Katherine Ching.